Accurate proteome-wide missense variant effect prediction with AlphaMissense
Por um escritor misterioso
Descrição
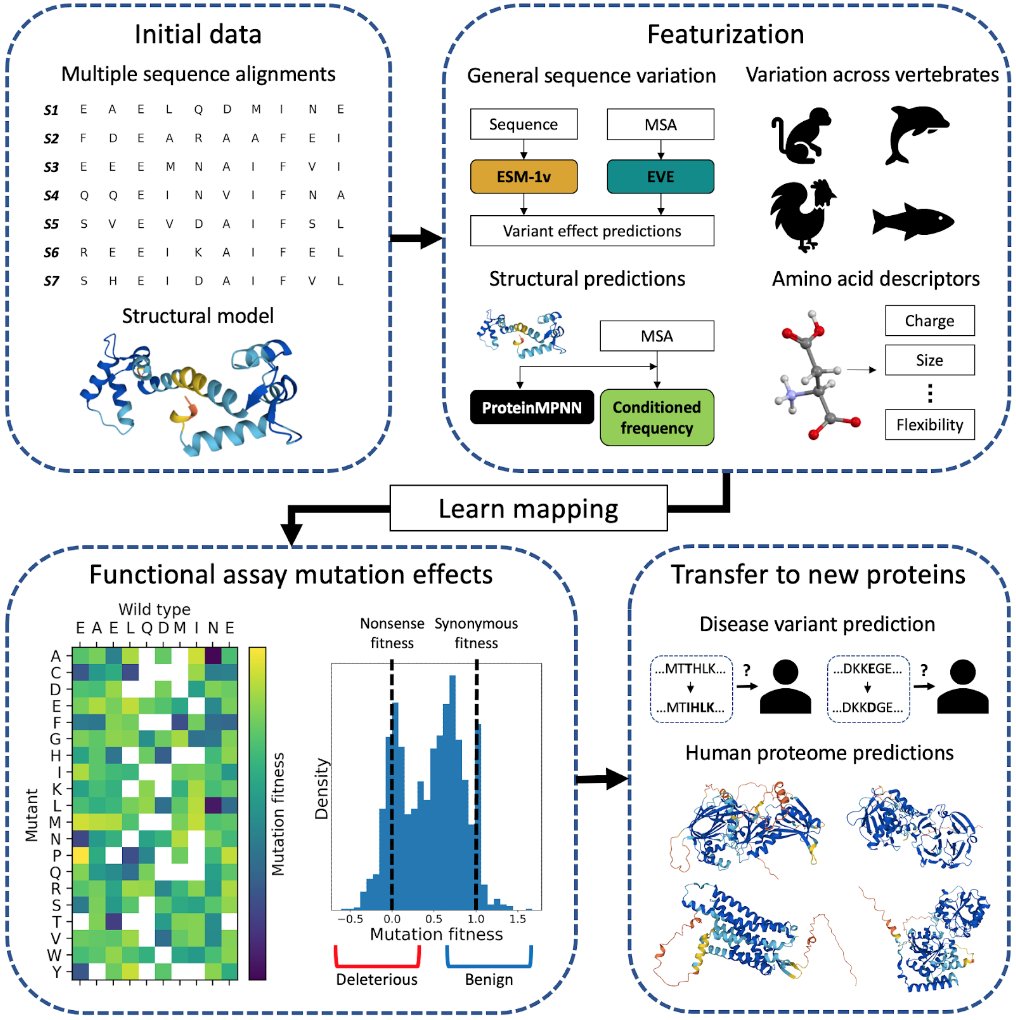
Yun S. Song on X: Predicting the effects of missense variants is a central problem in human genome interpretation. We are thrilled to share our preprint on using cross-protein transfer (CPT) learning

Evaluating DeepMind's AlphaMissense Classifier

Google DeepMind announces AI ``AlphaMissense'', which may help predict which genetic mutations are harmful and identify the cause of genetic diseases - GIGAZINE

PDF) Genome-wide prediction of disease variant effects with a deep protein language model

Accurate proteome-wide missense variant effect prediction with AlphaMissense

PDF) Genome-wide prediction of disease variant effects with a deep protein language model

Predicting the pathogenicity of missense variants using features derived from AlphaFold2

Predicting the pathogenicity of missense variants using features derived from AlphaFold2

The COSMIS score is more predictive of variant pathogenicity than other
A comprehensive resource of Bioinformatics and Genomics
de
por adulto (o preço varia de acordo com o tamanho do grupo)






/i.s3.glbimg.com/v1/AUTH_08fbf48bc0524877943fe86e43087e7a/internal_photos/bs/2022/t/c/0c2a7UQEGOYFGg3dhTxQ/minecraft-the-wild-update-atualizacao-warden.jpg)