Combining Machine Learning and Molecular Dynamics to Predict P-Glycoprotein Substrates
Por um escritor misterioso
Descrição
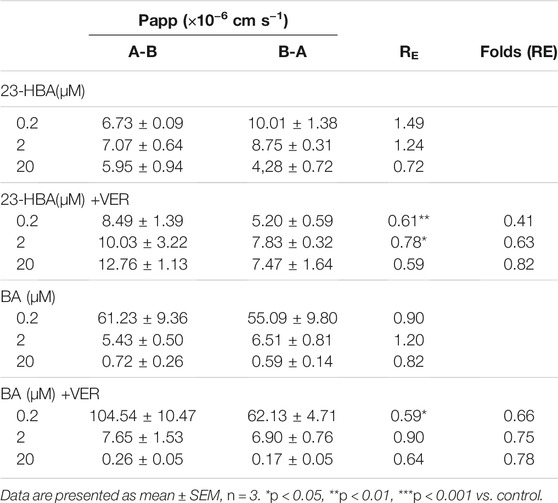
Frontiers Involvement of P-gp on Reversing Multidrug Resistance Effects of 23-Hydroxybetulinic Acid on Chemotherapeutic Agents

Combining Machine Learning and Molecular Dynamics to Predict P-Glycoprotein Substrates

Combining Machine Learning and Molecular Dynamics to Predict P-Glycoprotein Substrates

Machine learning for small molecule drug discovery in academia and industry - ScienceDirect

Combining machine‐learning and molecular‐modeling methods for drug‐target affinity predictions - Perez‐Lopez - 2023 - WIREs Computational Molecular Science - Wiley Online Library
Molecules, Free Full-Text

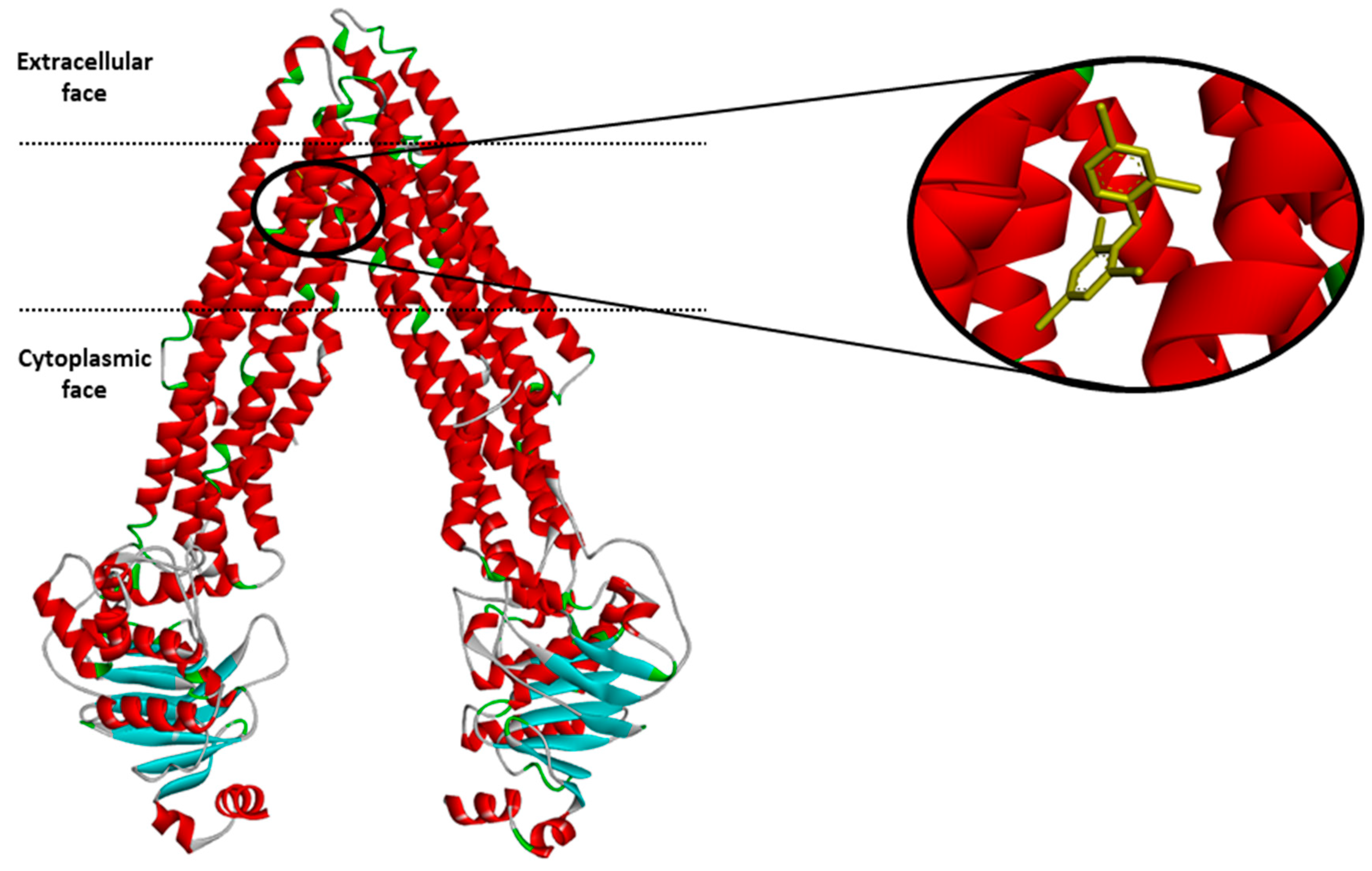
Molecular modeling of human P-gp structure. (a) 3D Structure of human

Combining Machine Learning and Molecular Dynamics to Predict P-Glycoprotein Substrates
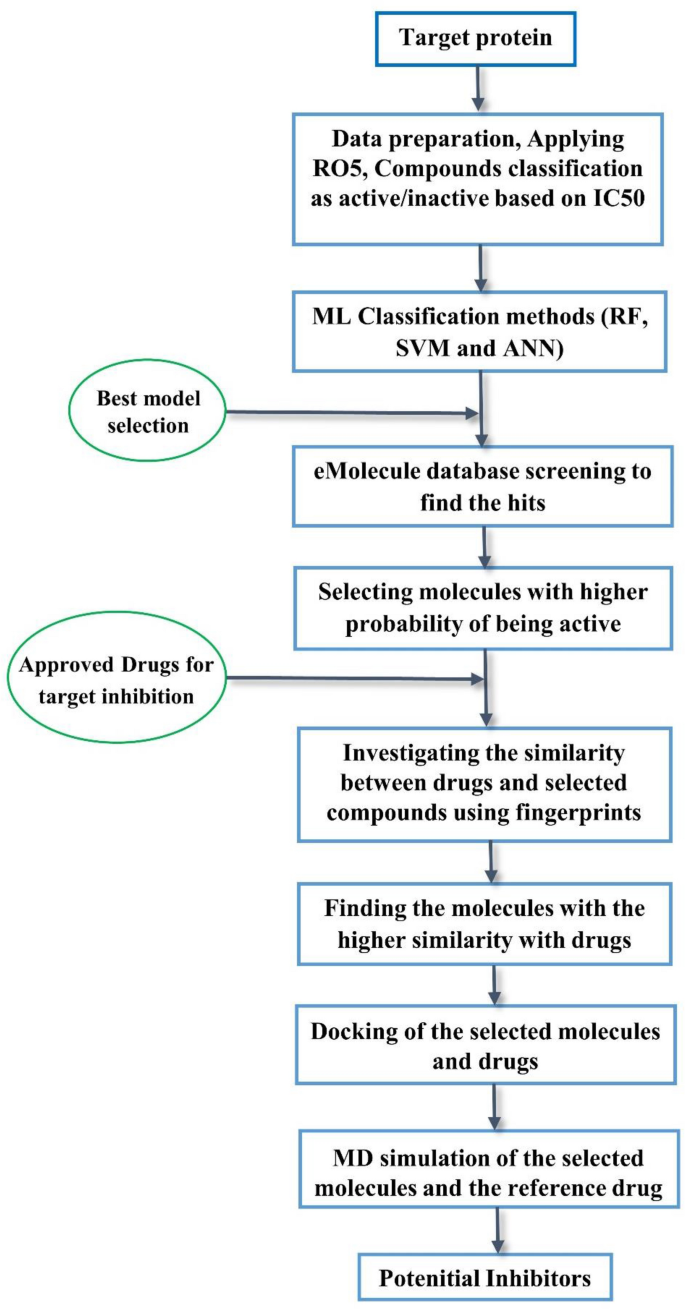
The use of machine learning modeling, virtual screening, molecular docking, and molecular dynamics simulations to identify potential VEGFR2 kinase inhibitors
de
por adulto (o preço varia de acordo com o tamanho do grupo)