cKBET: assessing goodness of batch effect correction for single-cell RNA-seq
Por um escritor misterioso
Descrição
lt;p>Single-cell RNA sequencing reveals the gene structure and gene expression status of a single cell, which can reflect the heterogeneity between cells. However, batch effects caused by non-biological factors may hinder data integration and downstream analysis. Although the batch effect can be evaluated by visualizing the data, which actually is subjective and inaccurate. In this work, we propose a quantitative method cKBET, which considers the batch and cell type information simultaneously. The cKBET method accesses batch effects by comparing the global and local fraction of cells of different batches in different cell types. We verify the performance of our cKBET method on simulated and real biological data sets. The experimental results show that our cKBET method is superior to existing methods in most cases. In general, our cKBET method can detect batch effect with either balanced or unbalanced cell types, and thus evaluate batch correction methods.</p>
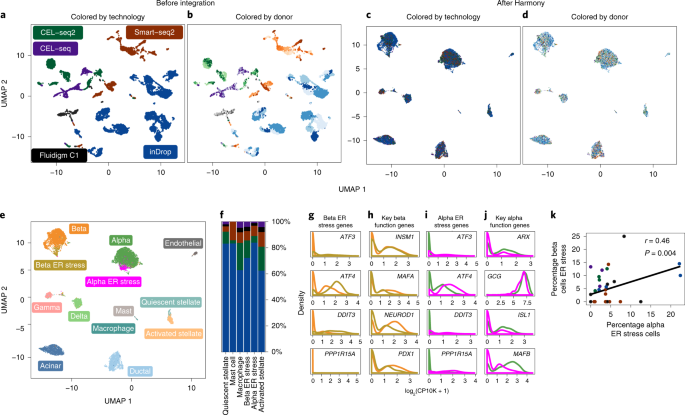
Fast, sensitive and accurate integration of single-cell data with Harmony

Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors

Assessment of batch-correction methods for scRNA-seq data with a new test metric
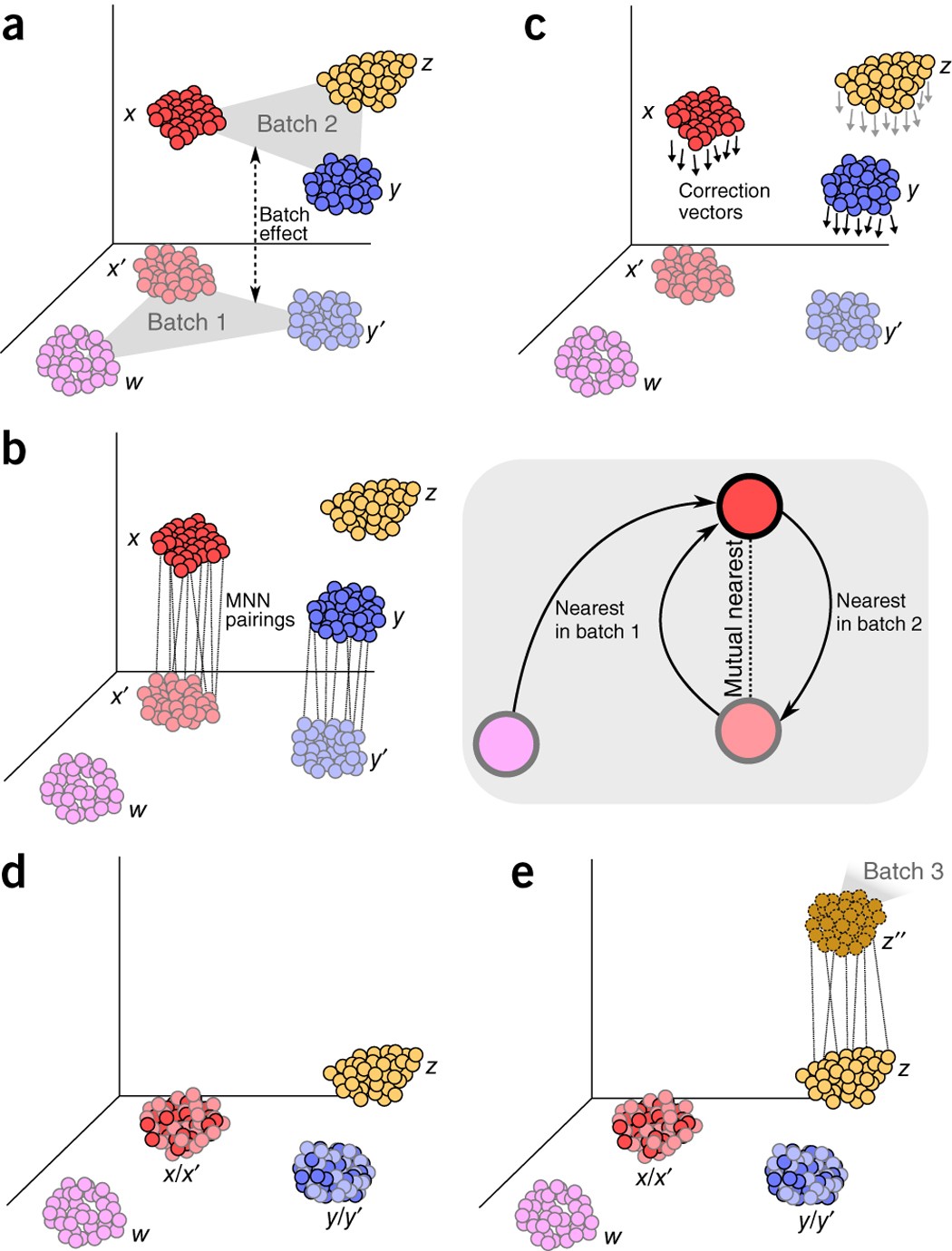
Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors

A comparison of methods accounting for batch effects in differential expression analysis of UMI count based single cell RNA sequencing - ScienceDirect
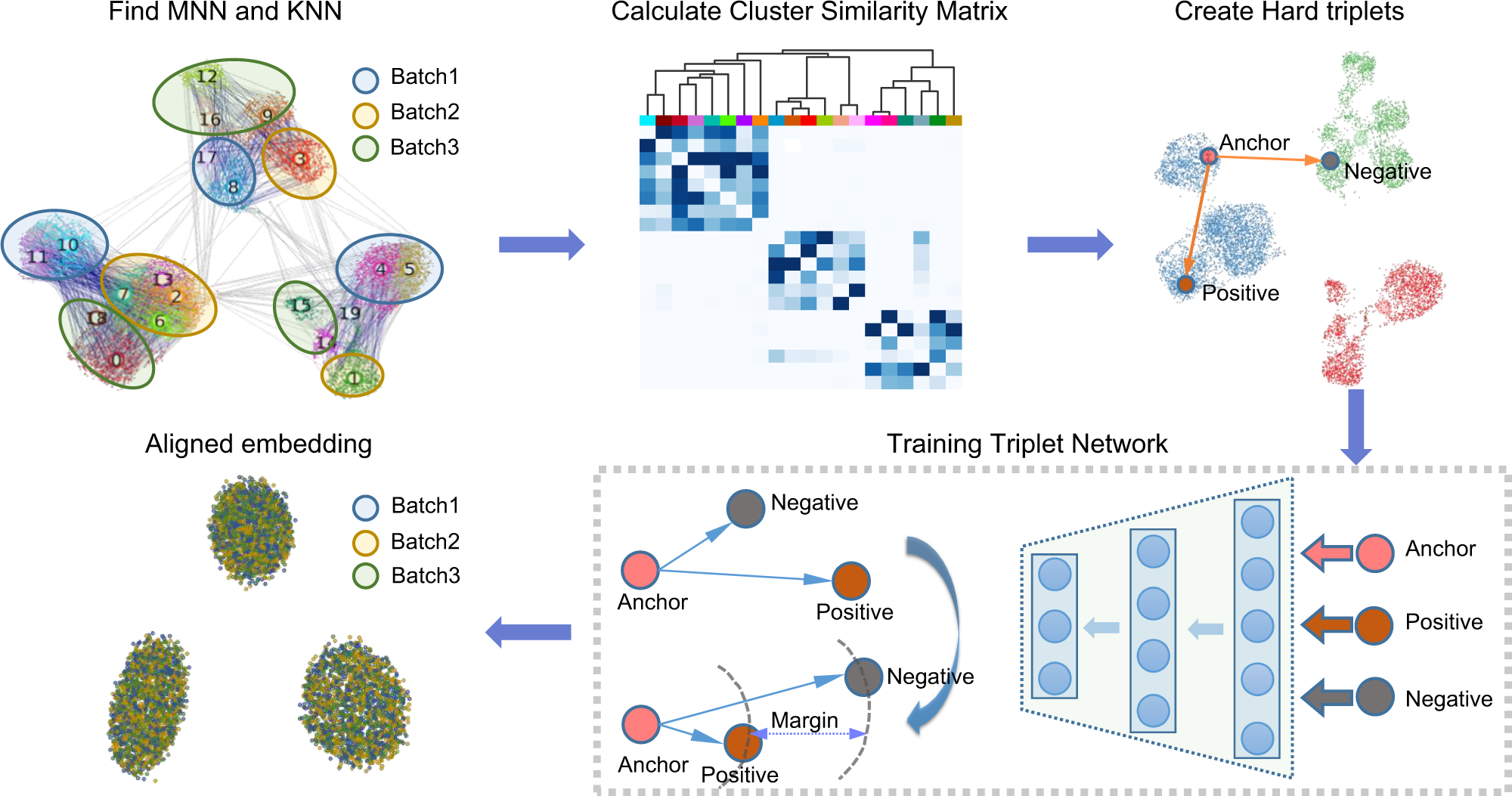
Batch alignment of single-cell transcriptomics data using deep metric learning
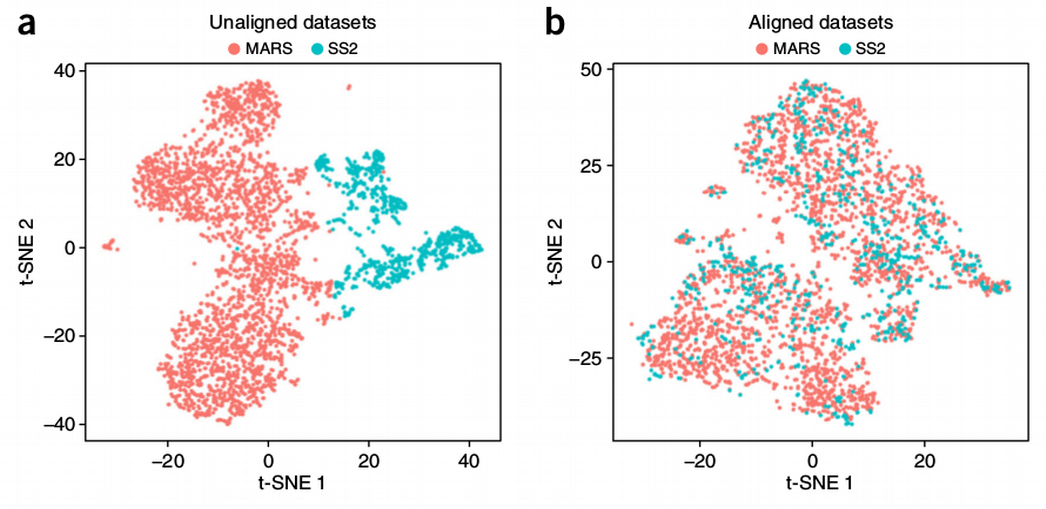
How to Batch Correct Single Cell. Comparing batch correction methods for…, by Nikolay Oskolkov

cKBET: assessing goodness of batch effect correction for single-cell RNA-seq
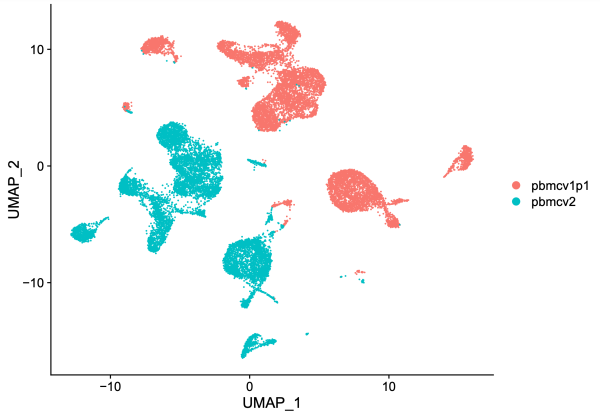
Batch Effect Correction in Chromium Single Cell ATAC Data - 10x Genomics
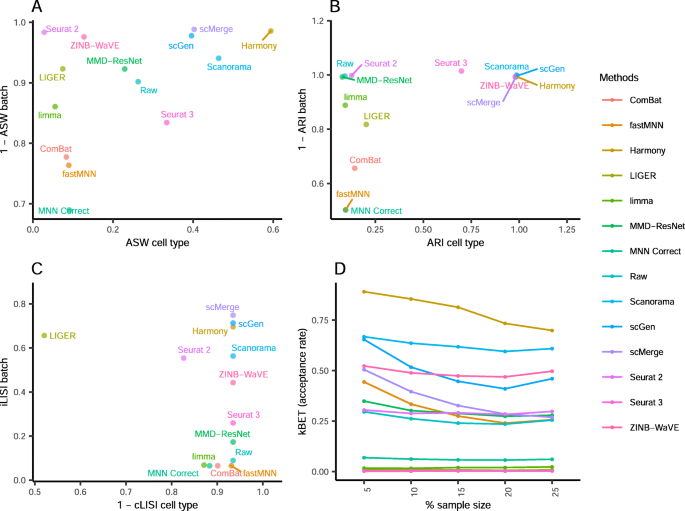
A benchmark of batch-effect correction methods for single-cell RNA sequencing data, Genome Biology
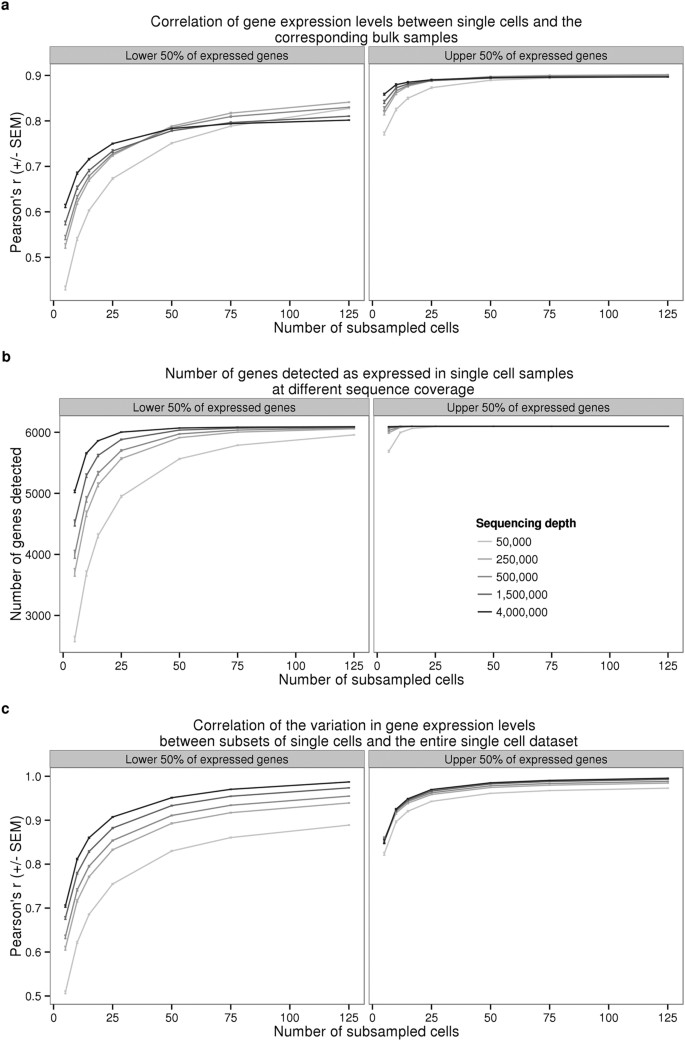
Batch effects and the effective design of single-cell gene expression studies

CellMixS: quantifying and visualizing batch effects in single-cell RNA-seq data
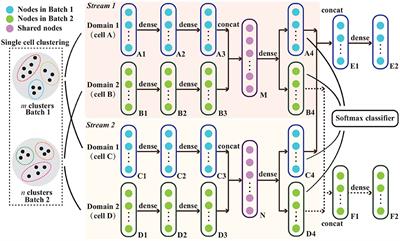
Frontiers CBA: Cluster-Guided Batch Alignment for Single Cell RNA-seq
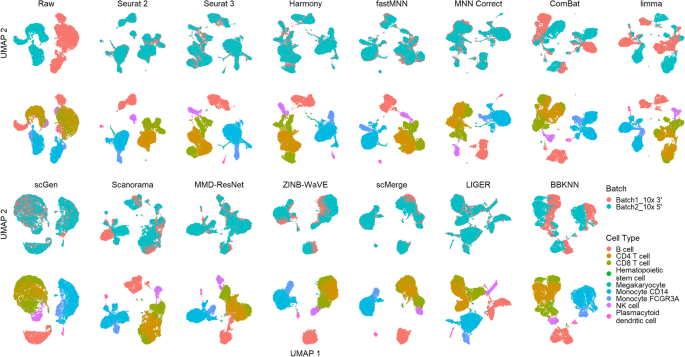
A benchmark of batch-effect correction methods for single-cell RNA sequencing data, Genome Biology
de
por adulto (o preço varia de acordo com o tamanho do grupo)