Modeling methyl-sensitive transcription factor motifs with an
Por um escritor misterioso
Descrição
Europe PMC is an archive of life sciences journal literature.

In need of good neighbours: transcription factors require local DNA hypomethylation for target binding
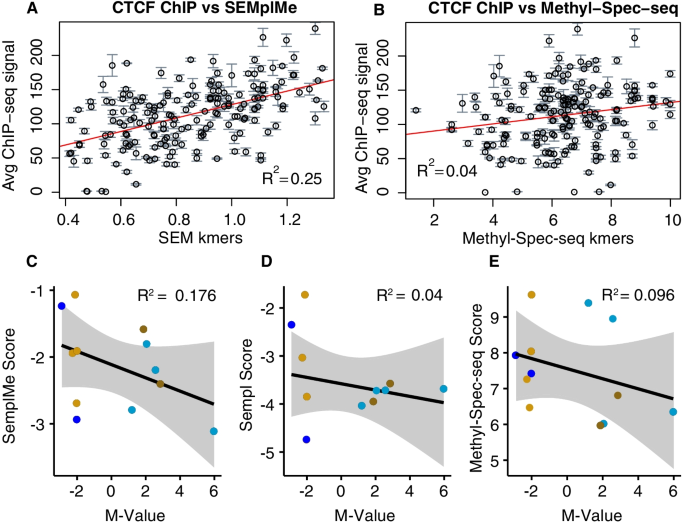
SEMplMe: a tool for integrating DNA methylation effects in transcription factor binding affinity predictions, BMC Bioinformatics
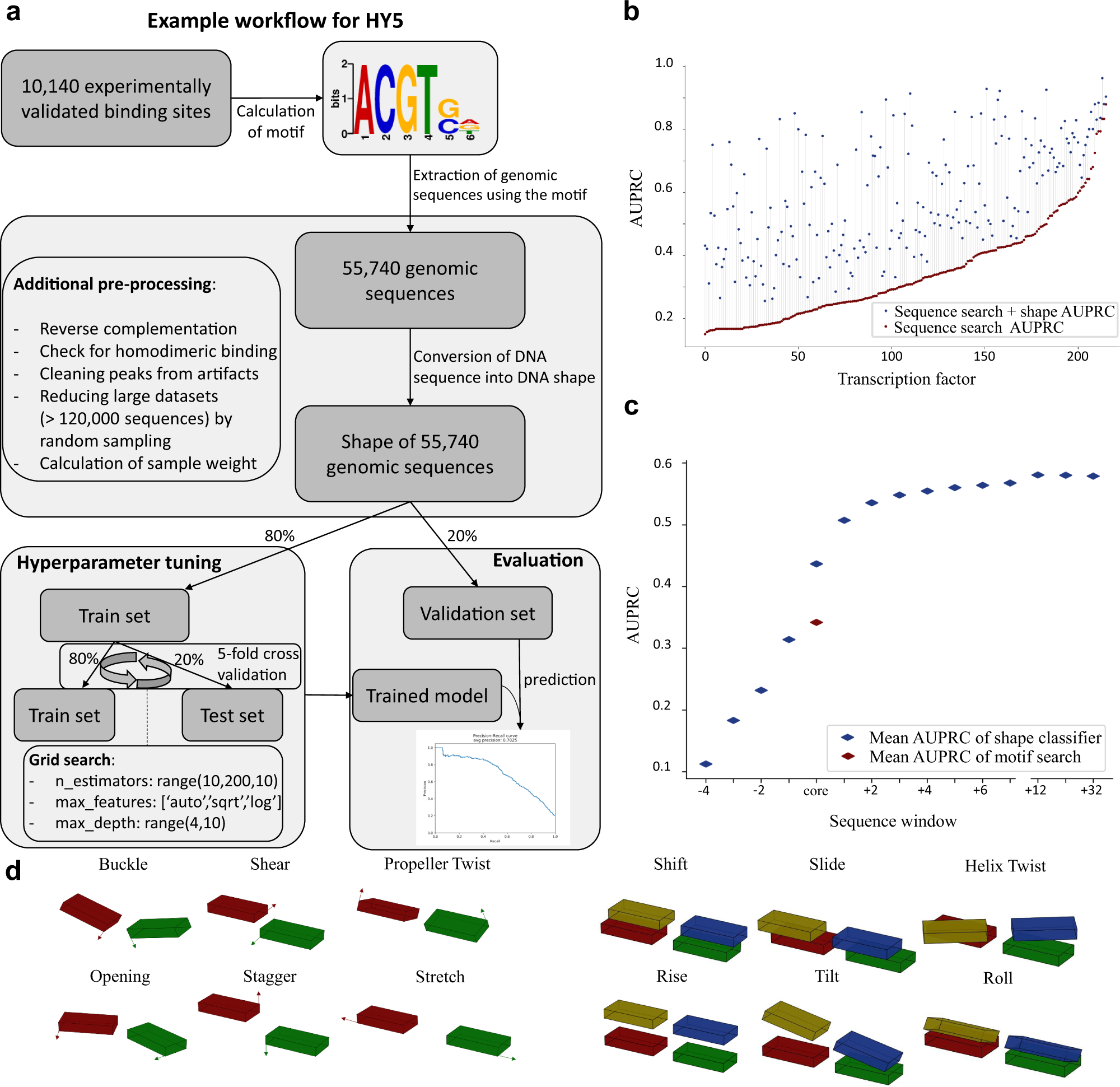
Local DNA shape is a general principle of transcription factor binding specificity in Arabidopsis thaliana
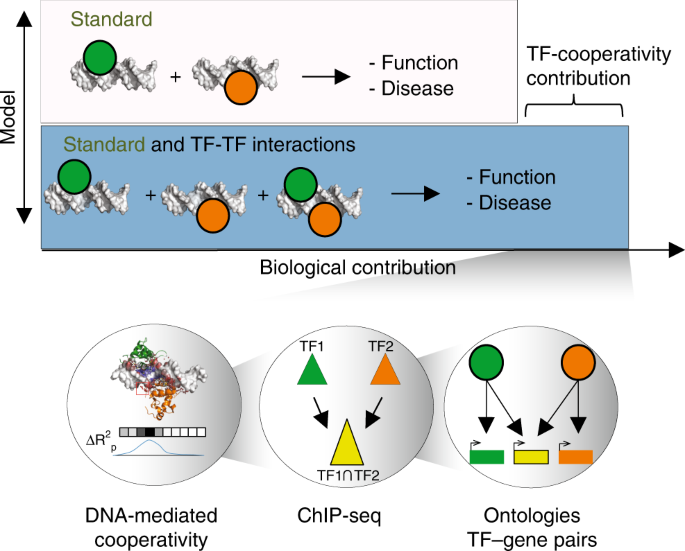
Mechanistic insights into transcription factor cooperativity and its impact on protein-phenotype interactions
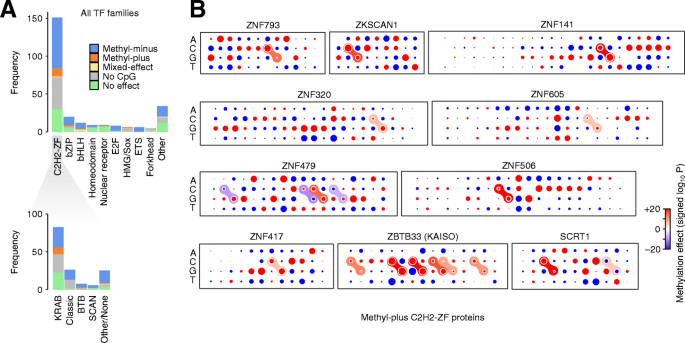
Toward a base-resolution panorama of the in vivo impact of cytosine methylation on transcription factor binding, Genome Biology

Predicting Preference of Transcription Factors for Methylated DNA Using Sequence Information - ScienceDirect

Quantitative Analysis of the DNA Methylation Sensitivity of Transcription Factor Complexes - ScienceDirect
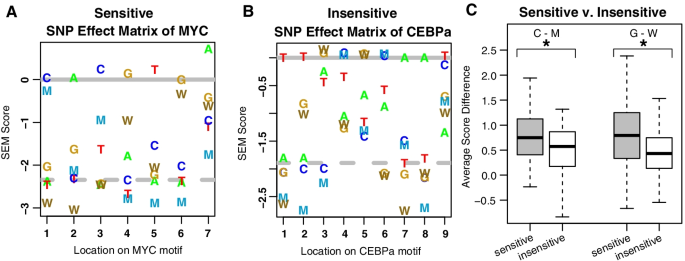
SEMplMe: a tool for integrating DNA methylation effects in transcription factor binding affinity predictions, BMC Bioinformatics

Noncanonical binding of transcription factors: time to revisit specificity?

DNA methylation - Wikipedia

Sensitivity of transcription factors to DNA methylation. - Abstract - Europe PMC
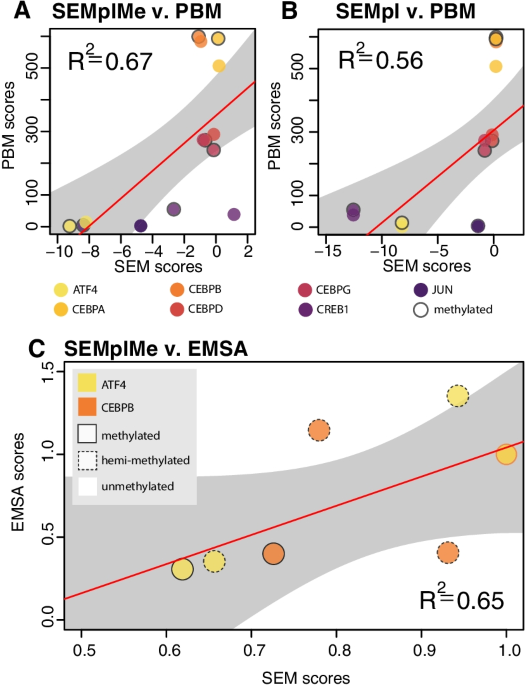
SEMplMe: a tool for integrating DNA methylation effects in transcription factor binding affinity predictions, BMC Bioinformatics

Modeling methyl-sensitive transcription factor motifs with an expanded epigenetic alphabet
Possible scenarios by which DNA methylation could impact TF binding (A)

Transcription factor binding site (TFBS) motif model. Sequences known
de
por adulto (o preço varia de acordo com o tamanho do grupo)







